Molecular Modeling Studies of Di-phenyl Alkyl Oxadiazoles, Sulfonamido-Phenyl Ethyl Ureas and Phenyl ethyl Amides for the Design of Novel Myosin Activators
Authors: Srihitha Akula1, Vani Madhuri Velavalapalli2, Sriharshith Akula4, Rambabu Gundla3*
1Department of Biology, Johns Hopkins University, Baltimore, Maryland, USA
2School of Pharmacy, Gitam University, Hyderabad, India
3Department of Chemistry, Gitam University, Hyderabad, India
4Herbert Wertheim College of Engineering, University of Florida, Gainesville, Florida, USA
*Corresponding author: Professor. Rambabu Gundla, Department of Chemistry, Gitam University, Hyderabad, India; E-mail: rgundla@gitam.edu
Received: 13 February 2023; Revision: 20 February 2023; Accepted: 23 February 2023; Published: 25 February 2023
Citation: Akula S, Velavalapalli VM, Akula S, Gundla G (2023) Molecular Modeling Studies of Di-phenyl Alkyl Oxadiazoles, Sulfonamido-Phenyl Ethyl Ureas and Phenyl ethyl Amides for the Design of Novel Myosin Activators 21st Century Cardiol, Volume 3 (1): 129
Abstract
Omecamtiv Mecarbil (OM) is a myosin activator that has been shown to combat reduced myocardial function. Recent studies conducted by Manoj Manickam et al. identified diphenylalkyloxadiazoles, sulfonamidophenylethylureas, and sulfonamidophenylethylamides as potential myosin activators. To further optimize, design and develop novel myosin activators, in the current study molecular docking studies (AutoDock (AD), AutoDock Vina (ADV)) and Quantitative Structure-Activity Relationship (QSAR) studies were conducted on these series of compounds. Our molecular docking studies have shown that a few structural modifications to the existing compounds that could lead to improved activity. Additionally, a QSAR equation was developed by correlating the molecular descriptors against the activity also a LUA based code was used for predicting the activity of the test set structures.
Keywords:
Omecamtive Mecarbil, Myosin activators, Molecular modeling, Molecular docking, AutoDock, AutoDockVina, Molecular descriptors, QSAR
Keywords: Omecamtive Mecarbil, Myosin activators, Molecular modeling, Molecular docking, AutoDock, AutoDockVina, Molecular descriptors, QSAR
Introduction
Heart failure (HF) affects more than 26 million people globally. Specifically, heart failure with reduced ejection fraction (Systolic Heart Failure) represents 50% of these cases [1,2]. Systolic heart failure can be attributed to impairment in left myocardial function and is identified when the ejection fraction is 40% or less [3]. Cardiac myosin is responsible for the contraction of myofilaments in the left ventricle and thus the pumping of blood in the heart [4]. In recent years, research has focused on developing and testing cardiac myosin activators such as Omecamtiv Mecarbil (OM) to combat reduced myocardial function [5,6]. These activators can bind to the active site on myosin and increase Left Ventricular Ejection Fraction (LVEF) [7]. Cardiac myosin goes through a multi-step process in which it contracts and relaxes to pump blood through the heart. At the initial stage, the myosin exists in a weakly bound state to the actin filament, and later it makes a strong bridge with actin filament [8]. In order to form, this bridge, myosin must be in a high-energy confirmation, which it achieves through ATP hydrolysis. OM targets and activates the myosin ATPase activity, thus triggering ATP hydrolysis and bringing myosin into a high-energy confirmation [9]. The formation of a strong bridge between actin and myosin is the rate-determining step in this process. The next step of this process is the power stroke, in which the actin-myosin complex release ADP and inorganic phosphate and cause the sarcomere to contract [10]. Targeting myosin ATPase allows the contraction state to be prolonged, causing the ventricle to fill up with more blood. [11]. OM thus increases ventricular systolic ejection time and fraction, giving it the capability to act against systolic heart failure [12]. Current studies on myosin activators have focused on OM as a reliable agent. Recent studies conducted by Manoj Manickam et al. have led to the discovery of three new class of compounds as cardiac myosin activators. These groups are diphenylalkyloxadiazoles, sulfonamidophenylethylureas and sulfonamidophenylethylamides [13,14,15]. In the current study, we used the above series of the molecular structures for molecular docking & QSAR studies. The molecular docking studies had been performed using AutoDock (AD) & AutoDock Vina ADV software’s [16,17]. From the molecular docking analysis, we found the critical interaction points at the active site by comparing the above reported high, low and moderately active compounds with OM. In addition to this, Quantitative structure-activity relationship (QSAR) studies were conducted against the biological activities reported in earlier studies. By mapping the relationship between molecular descriptors and biological activity, we were able to build a QSAR equation. By using this equation, we were able to discover the specific properties of a compound that make it active in the myosin binding site. A LUA based code was used to build a program for predicting the activity of newly designed compound, or a known test set structure. The results of our study allow us to define and optimize the properties of an ideal cardiac myosin activator. This will be further used to build novel compounds that will work as the best activators and therefore offer a resolution to the growing HF pandemic.
Materials & Methods
A total of one hundred compounds (Di-phenyl alkyl oxadiazoles, Sulfonamido phenyl ethylureas & Sulfonamido phenyl ethylamides) were built using Marvin Sketch software, and an open-source Babel program was used to convert them into 3D pdbqt file format for docking study [18,19]. The 3D crystal structure of the myosin activator was retrieved from the Protein Data Bank (PDB ID-5N69) [20]. AutoDock & AutoDock Vina software’s [16,17] were used for molecular docking studies. The receptor protein structure was prepared by removing OM, water, hetero & other metal atoms from the reported PDB dimer complex. Then the dimer complex was sliced into monomers for molecular docking. The polar hydrogens and Kollman charges were used for the docking process, and Gasteiger charges were added for the ligand preparation [21,22]. The grid box size for search space at the active site is defined as 40, 45 & 35 Å in x, y, z directions. To carry out the QSAR studies, the molecular descriptors were calculated using PaDEL software for each compound [23]. The dataset was divided into training and test sets. BuildQSAR 2.1.0 software [24] was used to calculate the correlation coefficient (R2), the adjusted r-square coefficient (R2adj), the standard error of estimate (SEE), the variance ratio (F) was used for evaluation of the obtained models [25]. Their quality was estimated by the square correlation coefficient (Q2) employing the leave-one-out cross-validation (LOO CV) method. Using online based LUA compiler (https://www.tutorialspoint.com/execute_lua_online.php) we developed approximately hundred lines of code for predicting the test set structures activities, and the procedure as follows.
BuildQSAR studies
The molecular structures were drawn by ACD/ChemSketch software (ACDLabs software package, version 12.0) [26]. The polarizability, superficial tension, volume, molar refractivity and log P along with 2318 molecular descriptors which including 1D, 2D and 3D. descriptors were calculated for each structure using PaDEL software [23]. In order to generate the high R2 we correlated molecular descriptors with the biological activities. These descriptors were used to build the QSAR equation using the BuildQSAR program [24]. Among the 2318 descriptors, we have chosen fourteen 2D descriptors that were showed the best correlation (Geary autocorrelation - lag 1 / weighted by mass, Sanderson electronegativities, first ionization potential, I-state. Normalized Randic-like eigenvector-based index from Barysz matrix / weighted by van der Waals volumes, topological distance matrix). The structural information in the form of the SMILES format of the 100 molecules, their calculated 2318 molecular descriptors and selected 14 descriptors is provided in excel spreadsheet format (Table S1 as a supplementary material).
LUA Methodology
The code uses three variables: 1) A string of all the names of the descriptors used, 2) A string of the QSAR formula, and 3) A string of the name, toxicity, and descriptor values of the tested compound. To begin, the first string is converted into a list of substrings (where each substring is the name of a descriptor) and compiled into a “new descriptors name” table. A similar process is done with the third string, but the values are converted from strings to numbers, creating a “new descriptor values” table. The “new descriptors name” table is then used to match the descriptor name with the descriptor values (in the “new descriptor values” table) for each descriptor and compile it into an array. The second string is then read through, and the uncertainty values are removed and replaced with an asterisk (multiplication operation). The second string is once again read through, however, this time the names of the descriptors are replaced with the values using reference from the previously compiled array. The formula string is then converted into an expression and processed to return the predicted value of the toxicity. This value is then compared with the original toxicity value given in the original third string and the residue and offset percent is calculated using simple arithmetic [23]. The developed LUA code is provided as supplementary material document (LUA Code for Test Set prediction).
Results & Discussion
Omecamtiv Mecarbil (OM) has been identified as a potential cardiac myosin activator and the compound is in the final stage of clinical studies [5,6]. Recently there have been three new classes of analogues namely, Di-phenyl alkyl oxadiazoles, Sulfonamido-phenyl ethyl ureas, and Phenyl ethyl amides reported as potential myosin activators. The current study is intended to identify the key molecular interactions between the reported activators and myosin activity. AutoDockVina (ADV) and AutoDock (AD) [16,17] molecular docking methods were used to identify potential molecular interactions of these reported activators that are critical for binding to myosin. In addition, Quantitative Structure Activity Relationships (QSAR) studies are performed to correlate the reported activities against their molecular descriptors of these molecular structures. The QSAR models are built to predict the activities of the newly designed small molecules.
Analysis of molecular docking interactions
The first step in the molecular docking study is the identification of the binding site and generation of the grid box around it. In this study, we used the OM bound myosin crystal structure (5n69) for the binding analysis and built multiple grids. In order to optimize the grid parameters, the OM structure was first docked using AD and ADV methods against multiple grid parameter boxes. The best grid box covering the binding site is found where the docked OM structure orientation is nearly the same as the reported crystal structure. This optimized grid box was used for docking of ninety-nine structures. OM specifically binds to the catalytic S1 domain of cardiac myosin, thereby stimulating myosin ATPase activity and improving energy utilization, when hydrolysis of ATP takes place. Consequently, the myosin-actin binding state becomes very strong, and the probability of the myosin heads bound to actin filaments increases, thus boosting ejection fraction or systolic function and reduced heart failures.
The OM-binding site is nestled in a narrow cleft separating two domains of the human cMD where it interacts with key residues that couple lever arm movement to the nucleotide state. OM binding induces an allosteric change in the conformation of three strands of a β-sheet that are crucial to communication between the nucleotide pocket and actin-binding interface. In the reported crystal structure, the bound OM is classified as four subunits based on the interactions with different sub domains of the myosin protein. (1) carboxymethyl-piperazine ring; - The OM is in an extended conformation with the carboxymethyl-piperazine group buried deeply in the cleft. (2) fluoro-benzine ring; - In bridging the gap, OM buries a bi-dentate salt bridge, E497-R712, beneath the fluoro-benzene ring. The fluoro-benzene ring of OM makes extensive packing interactions with residues along one face of the SH1 helix including C705 (SH1). The drug bridges the gap from the SH1 helix to the first strand (N711-I713) of the three-stranded β-sheet of the converter domain. (3) amino-carbamoyl linker; and (4) methyl-pyridinyl ring - the amido group of N711 forms a H-bond with the nitrogen of the OM methyl-pyridinyl ring. In the current analysis, a comparative analysis of high (oxa6a), moderate (spa21), & low active (ppu10) of the reported structures represented in Figure (Fig) 1 against OM was performed to find out the intermolecular interactions with binding site residues. Besides the influence of the conformational changes in the critical domains discussed above that are responsible for ATPase activity.
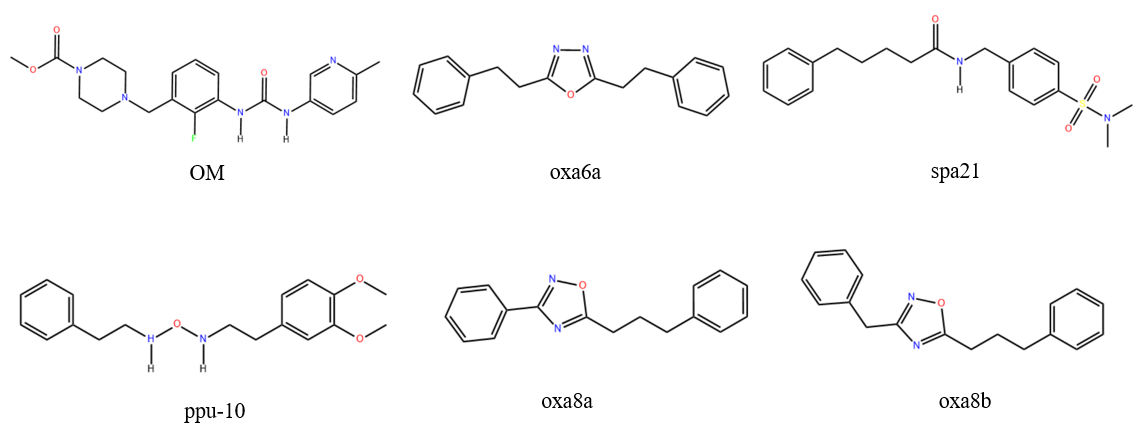
Figure 1: Structural information of Omecamtive Mecarbil (OM), reported myosin activators: diphenylalkyloxadiazoles (oxa6a, oxa8a, oxa8b), sulfonamidophenylethylureas (spa21) and sulfonamidophenylethylamide (ppu10).
Analysis of AutoDockVina (ADV)results
ADV docking scores were given in Table S1 as supplementary material. The binding orientations of OM, oxa6a, spa21 & ppu10 structures were represented in Fig.2. To understand the docking position of the reference structure, first we aligned with OM to compare the orientation with OM in the binding site (Fig.2a). We found that the carboxymethyl piperazine domain of OM is buried deep in the cleft of LYS-146 to SER-148 of alpha helix strand on one side, and with ALA-767 as well as GLY-771 to GLU-774 β-sheet on other side (Fig.2b). Docking results showed that the high and moderate activity molecules reside in this pocket, and their phenyl groups show π-alkyl bonding with the LEU-770 amino acid residue (Fig.2c-2d). On the other hand, the phenylpropylurea group of the low activity group ppu-10 faces away from this pocket and has carbon-hydrogen bond interactions with ASN-160 and LYS-145 (Fig.2e). The next domain we examined was the Fluoro benzene ring of the OM, which is present between the GLU-774, GLN-163, TYR-164 amino acids of alpha helix (Fig.2b). The high activity group oxa-6a formed π-anion interactions with the GLU-774 and TYR-164 groups (Fig.2c). The moderately active spa-21 formed π-π stacking interactions with the TYR-164 and pentamide of spa-21 formed H-bonding interactions with THR-167 (Fig.2d). The low activity compound Ppu-10, which does not have strong interactions with the TYR-164 residue (Fig.2e) and thus favorable interactions with the TYR-164 residue seemed to contribute to the activity. The amide group of methyl pyridinyl of OM is surrounded by the amino acids LEU-770, ASP-168, ARG-712, and ILE-713. The phenyl propyl group of the oxa-6a and phenyl pentamide of spa-21 are in the same pocket and form interactions with LEU-770 and ILE-713. On the contrary, the dimethoxy phenyl of ppu-10 is extended towards the HIS-666 and formed π-cation and H-bonding interactions with HIS -492 and GLU-497. The ADV results clearly indicate the differences between high(oxa6a), moderate (spa21) and low (ppu10) active compounds in terms of number of binding interactions at the myosin active site.

Figure 2: Figure 2: AutoDockVina (ADV) predicted bound conformations of (2a). Aligned OM, oxa6a, spa21, ppu10 (2b). OM(green), (2c).oxa-6a(blue), (2d).spa-21(pink), (2e).ppu-10(yellow) against 5N69 protein binding site. Analysis of AutoDock (AD) results
AD docking scores were given in Table S1 as supplementary material. The binding orientations of OM, oxa6a, spa21 & ppu10 structures were represented in Fig.3. The results obtained using AD were analyzed similarly as ADV, initially we aligned all the four structures (OM, oxa6a,spa21,ppu10) docked postures at the myosin active site (Fig.3a) for the comparative study. The carboxy methyl piperazine of OM is in the pocket of ASN-711, ARG-712 beta sheet on one side, ARG-722 and ASP-168, and TYR-164 alpha helix on other side (Fig.3b). Oxa-6a is slightly away from the pocket and one of the phenyl propyl groups is between the GLN-163 and GLU-774 and have π-π bond with TYR-164 (Fig.3c). The phenyl pentamide group of spa-21 is in the vicinity of ARG-712, ASN-711, LEU-770 and PHE-765, ARG-721 (Fig.3d) while the phenyl propyl group of ppu-10 is away from the OM pocket and towards the HIS -666, HIS- 492 (Fig.3e). The fluoro benzene ring of OM is present in the pocket of LEU-770, TYR-164, GLN-163 where the oxa-6a phenyl propyl group is also present. H-bonding interactions are observed between the hydroxy methyl pyridinyl amino group of OM and ARG-147 and ASN-160 in the pocket of ALA-767, LEU-120 Another phenyl propyl group of oxa-6a is also bound towards LEU-120, ALA-767 (Fig.3b-3c). There are H-bonding interactions observed between spa-21 phenyl pentamide and TYR-164, THR-167. The dimethyl sulfamoyl group of spa-21 is present in the same pocket of OM forming interactions with ASN-160 and LYS-145 (Fig.3d). Dimethoxy phenyl group of ppu-10 are in the same pocket forming interactions with TYR-164 and the phenyl group forms π-cation and π-anion interactions with ARG-721 and GLU-774 (Fig.3e). In both the AD and ADV results, the low active ppu-10 binding interactions at the active site are lower when compared with moderate and high active molecules, and it is shifted slightly from the pocket. This comparative study will give an intuition to design the novel myosin activators for developing novel drugs to control the HF disease.

Figure 3: AutoDock (AD) predicted bound conformations of (3a). Aligned OM, oxa6a, spa21, ppu10 (3b). OM(green), (3c). oxa-6a(blue), (3d). spa-21(yellow), (3e). ppu-10(pink) against 5N69 protein binding site.
Analysis of QSAR results
QSAR modeling is a type of regression modeling used to find and predict the correlation between structural properties (referred to as the descriptors) of a compound and its biological activity. QSAR is often employed in the fields of chemistry, biology and engineering and is a relatively established form of modeling. The process of QSAR involves first finding the relation between the properties and activity of compounds in the form of a “QSAR formula”. The formula is then used to predict the activity of new compounds [27]. The aim of this study is to correlate the molecular docking scores against the biological activities, so that the activities of newly designed compounds will be predicted based on the docking scores. A Training Set and TestSet are used to assess the accuracy of the derived formula. QSAR models were derived by multiple regression analyses that were performed using the BuildQSAR program [24] to determine the coefficients of the correlation equations. In all equations in this article, the numbers in parentheses represent the 95% confidence intervals of the coefficients, n is number of data points, r is the correlation coefficient, s is the standard deviation, q2 is the cross- validated value and F is Fisher value, which measures for the statistical significance. The multiple linear regression option allows the user to select any number of descriptors, without the involvement of subsets [28]. We used the MLR method to correlate the activities (dependent value) ‘vs’ independent variables (molecular docking scores) to find out the QSAR equation with highest correlation coefficient (R2) (Fig.4a-4b). Along with this, the systematic search feature can be used to generate statistical values for all possible combinations of descriptors. In our study, the R2 value (0.75) was found to be relatively satisfactory, whereas the adjusted R2 value was significantly lower (0.62). This indicates that although the QSAR model generated has a high R2 value, the addition of new terms to predict the activity, tend to decrease the reliability of its prediction. Similarly, the value of Q2 is much lower than the value of R2, showing that the predictability of the model is quite low. One thing to note however, is that the variance of the data as well as variance of the prediction, is very low, as shown by the SDEP and SPRESS statistics (Fig.4c). This makes the data spread very dense, therefore, predictions for activity outside of the data range would become highly inaccurate.
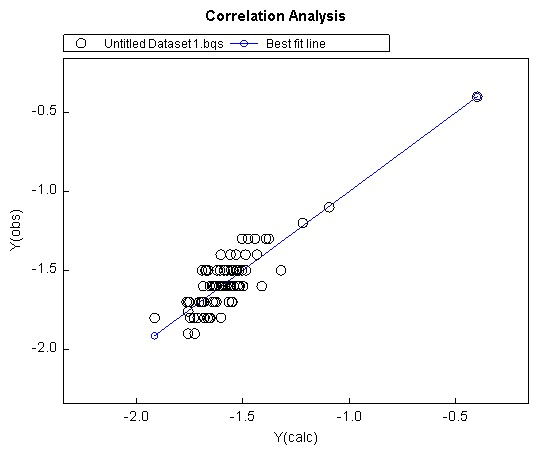
Figure 4a: QSAR correlation analysis.
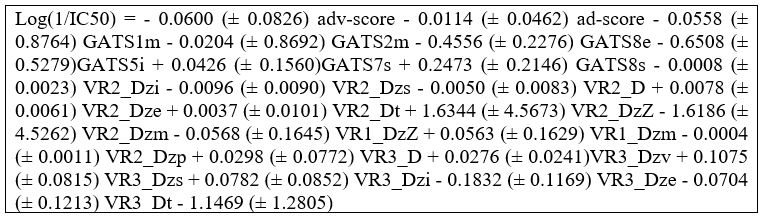
Figure 4b: QSAR Equation developed by using a Training set of 85 compounds.
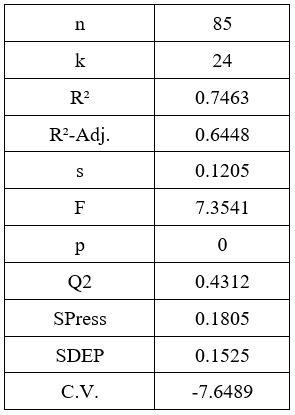
Figure 4c: QSAR Fitting Parameters Property Value.
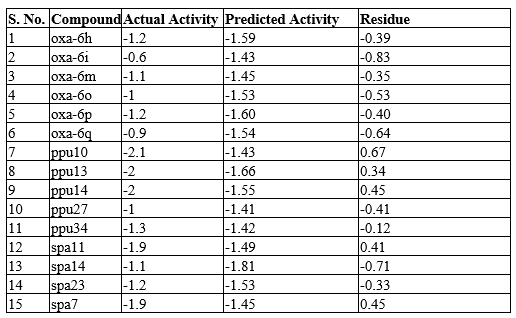
Table 1: Experimental activities (Actual activities) of 15 test set compounds are converted into -logIC50. The activities of the test set compounds are predicted using QSAR equation. Residual is difference between actual & predicted activities.
As mentioned previously the calculated molecular descriptors as given in supplementary material (Table S1). We found that the descriptors had the greatest influence on the IC50 values were the Geary autocorrelation descriptors and the Randic-like eigenvector-based indices. These descriptors are used to measure spatial autocorrelation, which is the correlation of a variable with itself through space (1) and a 2D weighted distance matrix accounting for the presence of heteroatoms and multiple bonds in the molecule, respectively. More specifically however, the most correlated descriptors were found to be in a 10-way tie between the following Randic-like eigenvector-based descriptors, with an R2 value of approximately 0.64. The descriptors with the prefix “VR2_D” (VR2_Dzi,VR2_Dzs,VR2_D,VR2_Dze,VR2_Dt,VR2_DzZ,VR2_Dzm,VR1_DzZ,VR1_Dzm,VR2_Dzp), represent normalized Randic-like eigenvector-based distances matrices. Normalization of the matrix causes it to account solely for the unit eigenvector, hence, the direction must play a larger role than the magnitude of the eigenvector. The descriptors with the prefix “VR1_D”, however, represent distance matrices which are not normalized, and the ones that showed high activity were in relation to the molecular weight of the compound (Tables S1).
The highest activity compound oxa-6a, moderately active compound spa21, and low activity compound ppu10 along with OM were all analyzed based on their structural elements (Figure 1). We observed that the content of nitrogen atoms and phenyl groups do not play a large role in influencing the activity. Rather, the content and positioning of oxygen appears to play a large role. In ppu10, there are three oxygen atoms, with a carbonyl group, and two ether groups, whereas in oxa-6a, there is only 1 oxygen atom in the center of the molecule. Like ppu10, OM contains three oxygen atoms, two of which are in a -NOOCH3 group, whereas spa21 contains two oxygen atoms in a sulfamoyl group. This seems to indicate that a greater content of oxygen contributes to a lower activity. The addition of sulfur and fluorine do not appear to have a noticeable influence on activity. In all three compounds, a group of nitrogen and oxygen can be seen at the center, along with two varied phenyl groups the ends of the compound, this is likely to be a condition that is required for initiation of ATP hydrolysis. Along with the composition, the symmetry of oxa-6a may also play a role in the activity of the compound. This can be affirmed by looking into low activity oxadiazoles molecules. Such examples include oxa-8a and oxa-8b (log1/IC50 = -1.8), which contain a very similar chemical composition and organization to oxa-6a, though, they are not symmetric, as shown in Figure 1. Looking at the training set (Given as supplementary data Table S2), it can be observed that the least accurate prediction was for the compound oxa-6a, likely due to its very high actual activity. In addition to this, inaccurate predictions were made for ppu10 and oxa-6q, with the actual activities of -2.1 and -0.9 respectively. Generally, it is difficult to predict very drastically high or low active compounds, whereas the compounds with activity around -1.3 have the most accurate predictions (Table 1).
Conclusion
In this study, novel di-phenyl alkyl oxadiazoles, sulfonamido phenyl ethylureas, and phenyl ethyl amides structures were used for molecular docking studies and QSAR analyses. Molecular docking results showed that π-π interactions between phenyl groups on the compounds with aromatic and neutral amino acid residues leads to high biological activity. π-cation interactions with basic residues such as Histidine and Arginine were correlated with low biological activity. The oxygen groups on the sulfonamido phenyl ethylureas and sulfonamido phenyl ethylamides interact unfavorably with basic residues. Symmetry in the compounds structure was also correlated with high activity. QSAR formula was generated to predict the biological activities of the compounds. The formula takes into account the structure of the molecules by specifically looking at several 2D molecular descriptors such as (Geary autocorrelation - lag 1 / weighted by mass, Sanderson electronegativities, first ionization potential, I-state. Normalized Randic-like eigenvector-based index from Barysz matrix / weighted by van der Waals volumes, topological distance matrix). A LUA based code developed and successfully applied for predicting the test set structures activity. Our modeling results will be useful for optimization patterns around of novel compounds and assess their capabilities for design potential myosin activators based on molecular docking interaction and their physico-chemical properties.
Disclosures
The authors have nothing to disclose.
Funding
No funding was involved in the completion of this review article.
References
1. Savarese G, Lund LH. Global public health burden of heart failure. Cardiac failure review. 2017 Apr;3(1):7-11. https://doi.org/10.15420/cfr.2016:25:2
2. Katz SD. Pathophysiology of chronic systolic heart failure. A view from the periphery. Annals of the American Thoracic Society. 2018 Feb;15(Supplement 1):S38-41. https://doi.org/10.1513/AnnalsATS.201710-789KV
3. University of Wisconsin Hospitals and Clinics Authority. “Heart Failure With Reduced Ejection Fraction (Systolic Heart Failure) | UW Health | Madison, WI.” UW Health, www.uwhealth.org/health/topic/special/heart-failure-with-reduced-ejection-fraction-systolic-heart-failure/tx4090abc.html
4. Inchingolo AV, Previs SB, Previs MJ, Warshaw DM, Kad NM. Revealing the mechanism of how cardiac myosin-binding protein C N-terminal fragments sensitize thin filaments for myosin binding. Proceedings of the National Academy of Sciences. 2019 Apr 2;116(14):6828-35. https://doi.org/10.1073/pnas.1816480116
5. Woody MS, Greenberg MJ, Barua B, Winkelmann DA, Goldman YE, Ostap EM. Positive cardiac inotrope omecamtiv mecarbil activates muscle despite suppressing the myosin working stroke. Nature communications. 2018 Sep 21;9(1):3838. https://doi.org/10.1038/s41467-018-06193-2
6. Teerlink JR. A novel approach to improve cardiac performance: cardiac myosin activators. Heart failure reviews. 2009 Dec;14:289-98. https://doi.org/10.1007/s10741-009-9135-0
7. Malik FI, Hartman JJ, Elias KA, Morgan BP, Rodriguez H, Brejc K, Anderson RL, Sueoka SH, Lee KH, Finer JT, Sakowicz R. Cardiac myosin activation: a potential therapeutic approach for systolic heart failure. Science. 2011 Mar 18;331(6023):1439-43. https://doi.org/10.1126/science.1200113
8. Andrei S, Iorgoveanu C. New perspective in heart failure management: could myosin activators be the answer?. Discoveries. 2014 Oct;2(4). https://doi.org/10.15190/d.2014.25
9. Kodera N, Ando T. The path to visualization of walking myosin V by high-speed atomic force microscopy. Biophysical reviews. 2014 Dec;6:237-60. https://doi.org/10.1007/s12551-014-0141-7
10. Sweeney HL, Hammers DW. Muscle contraction. Cold Spring Harb. Perspect. Biol. 2018; 10, a023200. https://doi.org/10.1101/cshperspect.a023200
11. Pollock JD, Makaryus AN. Physiology, cardiac cycle. InStatPearls [Internet] 2022 Oct 3. StatPearls Publishing.
12. Liu C, Kawana M, Song D, Ruppel KM, Spudich JA. Controlling load-dependent kinetics of β-cardiac myosin at the single-molecule level. Nature structural & molecular biology. 2018 Jun;25(6):505-14. https://doi.org/10.1038/s41594-018-0069-x
13. Manickam M, Jalani HB, Pillaiyar T, Boggu PR, Sharma N, Venkateswararao E, Lee YJ, Jeon ES, Son MJ, Woo SH, Jung SH. Design and synthesis of sulfonamidophenylethylureas as novel cardiac myosin activator. European Journal of Medicinal Chemistry. 2018 Jan 1;143:1869-87. https://doi.org/10.1016/j.ejmech.2017.10.077
14. Manickam M, Boggu PR, Pillaiyar T, Sharma N, Jalani HB, Venkateswararao E, Jung SH. Exploration of diphenylalkyloxadiazoles as novel cardiac myosin activator. Bioorganic & Medicinal Chemistry Letters. 2018 Aug 1;28(14):2369-74. https://doi.org/10.1016/j.bmcl.2018.06.031
15. Manickam M, Pillaiyar T, Namasivayam V, Boggu PR, Sharma N, Jalani HB, Venkateswararao E, Lee YJ, Jeon ES, Son MJ, Woo SH. Design and synthesis of sulfonamidophenylethylamides as novel cardiac myosin activator. Bioorganic & Medicinal Chemistry. 2019 Sep 15;27(18):4110-23. https://doi.org/10.1016/j.bmc.2019.07.041
16. Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. Journal of computational chemistry. 2009 Dec;30(16):2785-91. https://doi.org/10.1002/jcc.21256
17. Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. Journal of computational chemistry. 2009 Dec;30(16):2785-91. https://doi.org/10.1021/acs.jcim.1c00203
18. Marvin was used for drawing, displaying and characterizing chemical structures, substructures and reactions, Marvin 6.2.2, 2014, ChemAxon http://www.chemaxon.com
19. Open Babel development team. (2016). Open Babel. Retrieved from http://openbabel.org/wiki/Main_Page
20. Planelles-Herrero VJ, Hartman JJ, Robert-Paganin J, Malik FI, Houdusse A. Mechanistic and structural basis for activation of cardiac myosin force production by omecamtiv mecarbil. Nature communications. 2017 Aug 4;8(1):190. https://doi.org/10.1038/s41467-017-00176-5
21. Singh UC, Kollman PA. An approach to computing electrostatic charges for molecules. Journal of computational chemistry. 1984 Apr;5(2):129-45. https://doi.org/10.1002/jcc.540050204
22. Gasteiger J, Marsili M. Iterative partial equalization of orbital electronegativity—a rapid access to atomic charges. Tetrahedron. 1980 Jan 1;36(22):3219-28. https://doi.org/10.1016/0040-4020(80)80168-2
23. Allouche AR. Gabedit—A graphical user interface for computational chemistry softwares. Journal of computational chemistry. 2011 Jan 15;32(1):174-82. http://dx.doi.org/10.1002/jcc.21600
24. De Oliveira DB, Gaudio AC. BuildQSAR: a new computer program for QSAR analysis. Quantitative Structure?Activity Relationships: An International Journal Devoted to Fundamental and Practical Aspects of Electroanalysis. 2000 Dec;19(6):599-601. https://doi.org/10.1002/1521-3838(200012)19:6<599::AID-QSAR599>3.0.CO;2-B
25. Ierusalimschy R, De Figueiredo LH, Filho WC. Lua—an extensible extension language. Software: Practice and Experience. 1996 Jun;26(6):635-52. https://doi.org/10.1002/(SICI)1097-024X(199606)26:6<635::AID-SPE26>3.0.CO;2-P
26. ACD/ChemSketch, version 2021.1.2, Advanced Chemistry Development, Inc. (ACD/Labs), Toronto, ON, Canada, www.acdlabs.com
27. Muhammad U, Uzairu A, Ebuka Arthur D. Review on: quantitative structure activity relationship (QSAR) modeling. J Anal Pharm Res. 2018;7(2):240-2. https://doi.org/10.15406/japlr.2018.07.00232
28. Ambure P, Halder AK, Gonzalez Diaz H, Cordeiro MN. QSAR-Co: An open source software for developing robust multitasking or multitarget classification-based QSAR models. Journal of chemical information and modeling. 2019 May 14;59(6):2538-44. https://doi.org/10.1021/acs.jcim.9b00295